|
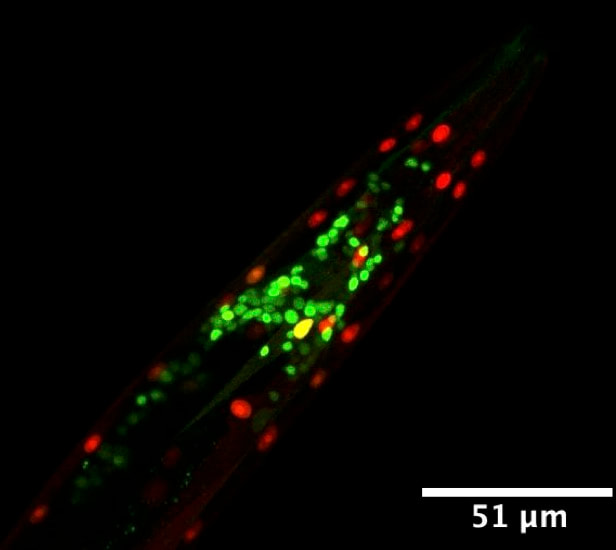
We have been using the tiny nematode worm Caenorhabditis elegans to uncover principles and mechanisms underlying tissue- and neuron subtype-specific alternative splicing patterns. We utilize a variety of approaches, including forward genetic screens with two-colour splicing reporters (diagram to the right) to identify novel regulators of splicing. More recently, we have utilized mRNA profiling strategies to measure splicing patterns across the transcriptome of various cell and tissue types. Our work is starting to uncover stereotyped regulatory outcomes in single neuron types within the nervous system, revealing a remarkable degree of complexity at the level of splicing regulation, even in an organism with a compact nervous system.
|